WaveletSEG
|
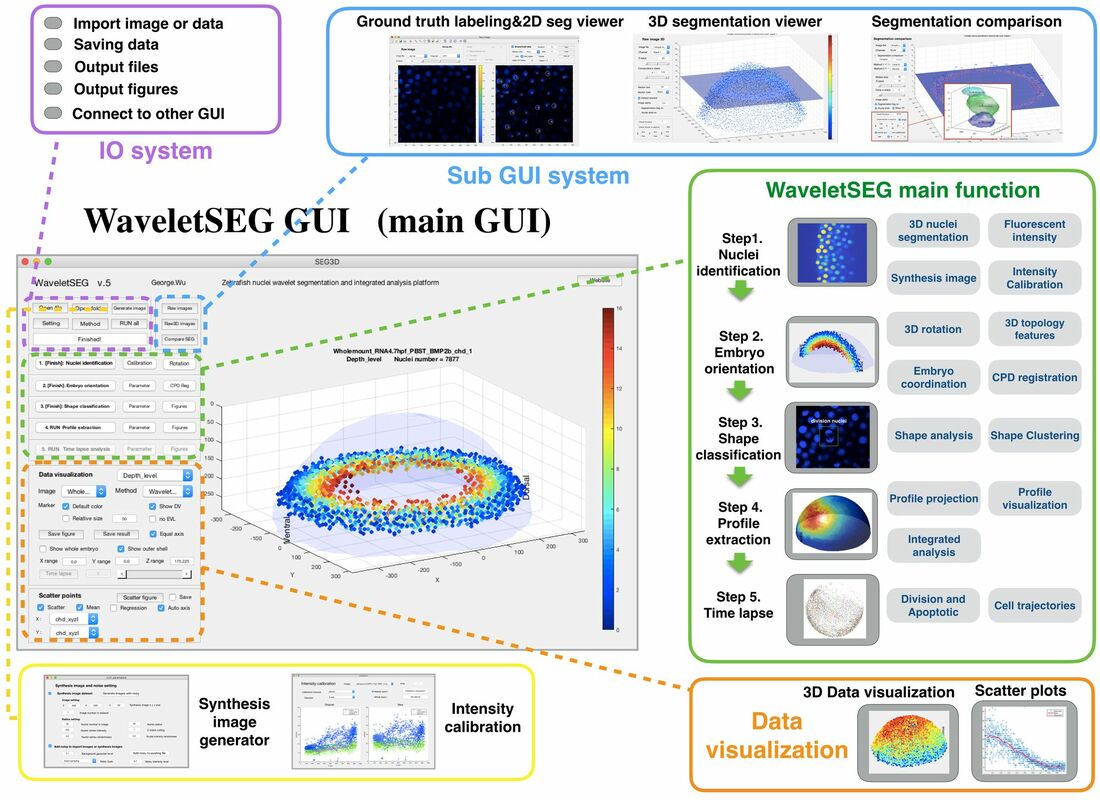
We develop novel wavelet-based segmentation algorithm which can do robust nuclei segmentation or single-molecule RNA segmentation without any prepossessing or thresholding steps, and separate overlapping cells accurately. We incorporate wavelet based nuclei identification, image registration, 3D topology features, and shape classification in Zebrafish whole mount embryo in automatically analysis platform named WaveletSEG. Three sub GUI are also created for segmentation results validation and methods comparison.
Tzu-Ching Wu, David Umulis, A novel automatically wavelet-based 3D nuclei segmentation method in Embryonic imaging. (in process) |
ZebEmbIM
|
To perform single cell transcriptome analysis of Zebrafish whole mount embryo, we created comprehensive analysis system termed ZebEmbIM. ZebEmbIM contains wavelet-based nuclei segmentation and single-molecule RNA segmentation. We also developed new membrane segmentation method and machine learning based Nascent RNA detection in ZebEmbIM.
Tzu-Ching Wu, David Umulis, Comprehensive Zebrafish embryonic imaging analysis system combined nuclei, RNA, and membrane segmentation using wavelet-based method. (in process) |
Lobefinder
|
We developed a convex-hull based algorithm termed LobeFinder to identify lobes, quantify geometric properties, and create a useful graphical output for further analysis. The ability to objectively count and detect new lobe initiation events provides an improved quantitative framework to analyze mutant phenotypes, detect symmetry-breaking events in time-lapse image data, and quantify the time-dependent correlation between cell shape change and intracellular factors that may play a role in the morphogenesis process. Tzu-Ching Wu, Samuel Belteton, Jessica Pack, David Umulis, Daniel B. Szymanski, Quantitative image analysis of pavement cell morphogenesis with LobeFinder. Plant Physiology, 171.4 (2016): 2331-2342. (* Selected as cover article) |
CCRIT
|
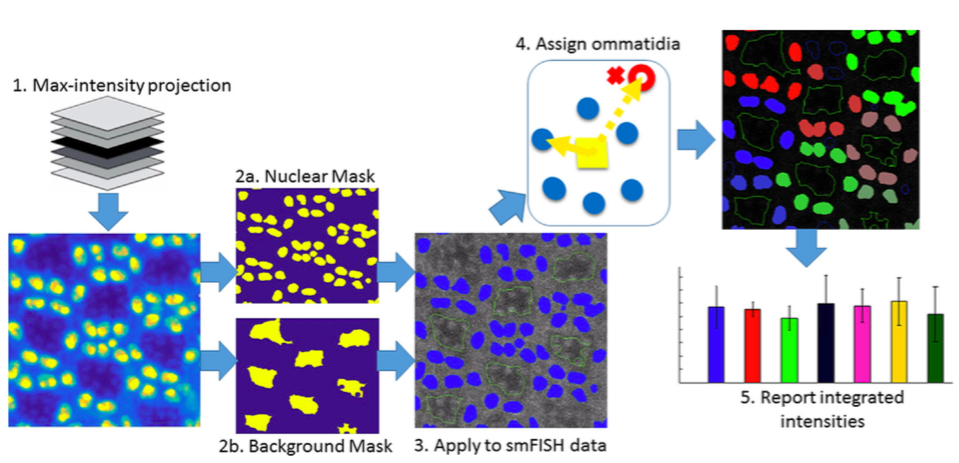
A noisy image of fluorescently-labeled mRNA transcripts can be analyzed by Cell-by-Cell Relative Integrated Transcript (CCRIT) Quantification to automatically identify cells and cell clusters and quantify each cell’s mRNA expression level. Pharris, M. C., Wu, T. C., Chen, X., Wang, X., Umulis, D. M., Weake, V. M., & Kinzer- Ursem, T. L. (2017). An automated workflow for quantifying RNA transcripts in individual cells in large data-sets. MethodsX. (SCI) |
Neuronal CalciumIM
|
We are developing integrated imaging analysis tool for two-photon calcium imaging in Zebrafish optic tectum. The spatio-temporal distribution of neurons population can be calculated and clustered after applying continuous wavelet transform on time series signal.
|
Cytoneme Simulation
|
We present the first cytoneme-mediated morphogen transport model to simulate the transport of Decapentaplegic (Dpp) in the imaginal wing disc epithelium of Drosophila. Cytoneme-mediated transport is viewed as a stochastic process effected by morphogen concentration distribution and cytoneme length between morphogen-producing cell and morphogen-receiving cell. Tzu-Ching Wu, David Umulis, A novel cytoneme-mediated morphogen transport model to establish Dpp gradient in Drosophila wing imaginal disc. (in process) |
Other imaging softwareI also developed several image processing codes with different purposes such as image tiling, fluorescent intensity, primary cilia segmentation with combining with machine learning, fuzzy logic, and optimization methods.
|
|
Cubix Health Technologies |
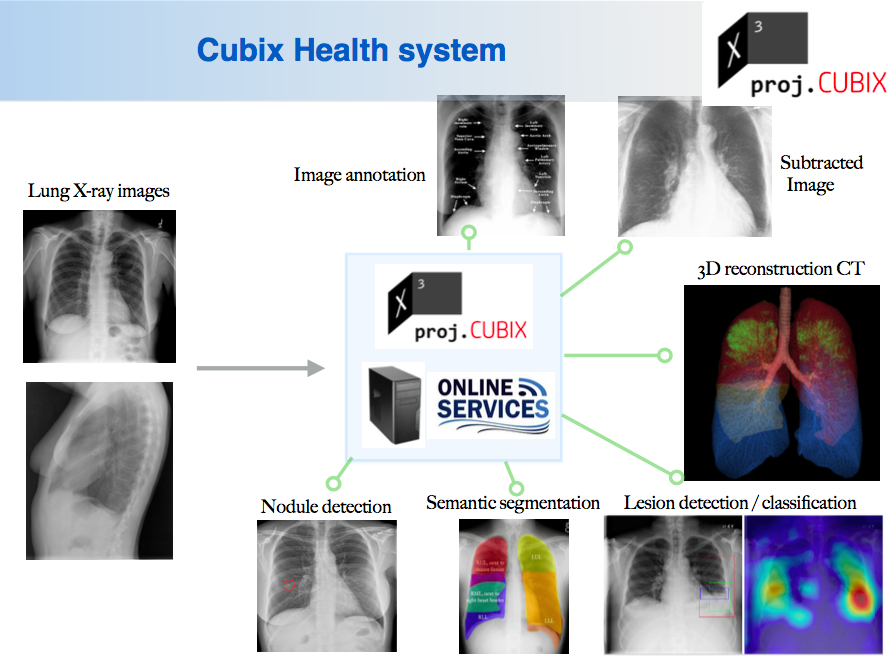
Cubix Health Applications (Cubix) develop innovative software package capable of revitalizing the use of affordable chest X rays in lung cancer screening. Our software process X-rays obtained from multiple angles and reconstruct a 3D model of the patient's chest marked with identified lung nodules
|